Kingston's genome sequencing research is contributing to fight against antimicrobial resistance and the development of novel probiotics
Bacterial infections are a major cause of deaths and illnesses across the world. The rise of antibiotic resistance in bacteria is a growing health concern, which has recently been widely highlighted in the media, and developing alternatives to common antibiotics is a government research priority.
Fundamental genomics studies at Kingston University are providing keys to the origin and evolution of bacterial species, and making a significant contribution to the development of novel antibacterials.
Deciphering genomes and discovering new bacteria
A series of fundamental genomics studies conducted by a team of microbiologists led by Andrey Karlyshev, Professor of Microbiology at Kingston University, has resulted in the deciphering of bacterial genomes of various bacterial species, including pathogens, probiotics and environmental isolates.

One set of experiments allowed the identification of bacteria preserved for millions of years in the permafrost of northern Russia. The identified bacterial species have never been described before, and provide a snapshot of the microbial environment on Earth many years ago. It also allows the researchers to investigate the evolution and the origin of bacteria. The results of these studies are being reported at the conference "Life in the Universe: physical, chemical and biological aspects" (June 2016, Moscow).
Another series of studies was devoted to the investigation of such important pathogens as Campylobacter jejuni and Neisseria meningitidis. Deciphering of the genomes of various strains of these species allowed the identification of traits responsible for antibiotic resistance and virulence, thus assisting in the development of novel antibacterials.
Sequencing of a number of genomes of beneficial (probiotic) bacteria was another direction of studies conducted by researchers led by Karlyshev. In addition to lactic acid producing bacteria, (Lactobacillus spp etc.), the genomes of Corynebacterium diphtheriticum, Enterococcus spp. have been sequenced. These studies identified specific genomic regions responsible for probiotic activities.
Production of complete genome sequence
One remarkable achievement was the deciphering of a complete genome sequence of a particular probiotic strain of Lactobacillus fermentum. The sequence of the genome consisting of a chromosome and plasmid (Fig1, A and B) was derived using Ion Torrent PGM, OpGen optical mapping, PacBio sequencing and a wide range of bioinformatics tools.
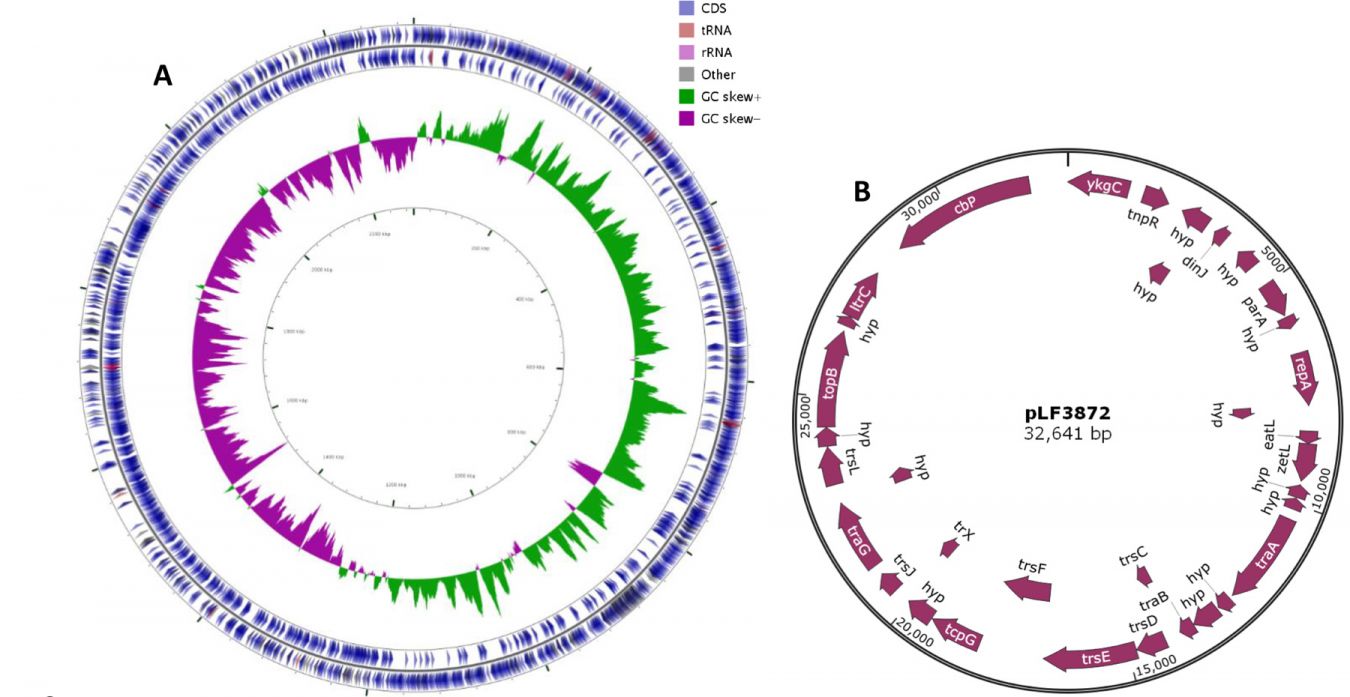
Fig. 1. Graphical image of a complete sequence L. fermentum genome represented by a chromosome (A) and plasmid (B)
This is one of few studies in the world on the production of a complete genome sequence (as opposed to draft genomes), and it is the first completed bacterial genome sequence produced by Kingston University.
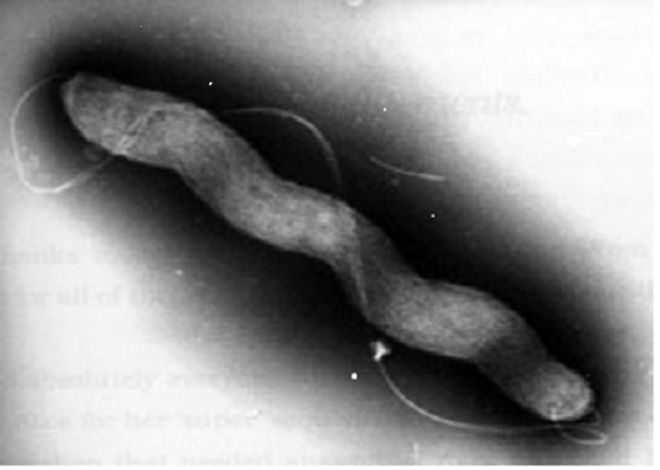
Fig. 2. Electron micrograph of Campylobacter jejuni showing a typical spiral shape of the cell and a long flagellum required for bacterial motility
This work also allowed the identification of specific features of probiotic bacteria that can be used for fighting gastrointestinal infections caused by Campylobacter jejuni (Fig.2), which is the most important gastrointestinal pathogen.
The recent in vitro studies conducted in Karlyshev's lab produced very promising results, which have recently been reported at the Microbiological Society Conference in Liverpool (April 2016).
Karlyshev's studies in the area of molecular microbiology have been published in a number of high ranking journals. He co-authored two ground-breaking research articles in Nature reporting the first completed genome sequences of human pathogens Campylobacter jejuni and Yersinia pestis (a causative agent of plague).
Karlyshev said that, despite recent revolutionary developments in the genome sequencing technology, the production of complete genome sequences remained a challenging task.
"One example is a human genome. Despite that it was reported many years ago, the human genome sequence remains incomplete even now. The vast majority of published bacterial genomes are also reported as 'draft' or incomplete sequences. Understanding what makes one probiotic strain much more efficient than another in fighting bacterial pathogens and infection, is important for the development of novel probiotics. Comparative genomics allows the identification of specific features of beneficial strains. The usage of probiotics has recently attracted huge interest, as these bacteria are naturally present in a human body. The risks of side effects and development of resistant pathogenic bacteria during treatment with probiotics are also significantly lower compared with those associated with the use of common antibiotics."
Contact us
- For non-student research enquiries, email the Research Office
- Research contacts
- Graduate Research School contacts
- Business and Enterprise contacts
- How to get to Kingston University
Contact us
- For non-student research enquiries, email the Research Office
- Research contacts
- Graduate Research School contacts
- Business and Enterprise contacts
- How to get to Kingston University


